Initially the "SSCP" method (for: single-strand conformation polymorphism) was developed in order to detect point mutations; which is the alteration of a single base within a longer sequence. Adapted to so-called "phylogenetic markers", like e.g the 16S rRNA gene for bacteria or the 18S rRNA for fungi, respectively, where different sequences relate to different microorganisms, this technique of sequence-specific separation of DNA allows to identify as well as to quantify all members of a microbial community.
AMODIA has extended this technique in a way that now even functional genes may be used for such studies.
The Universal Molecular Identification at AMODIA follows this scheme:
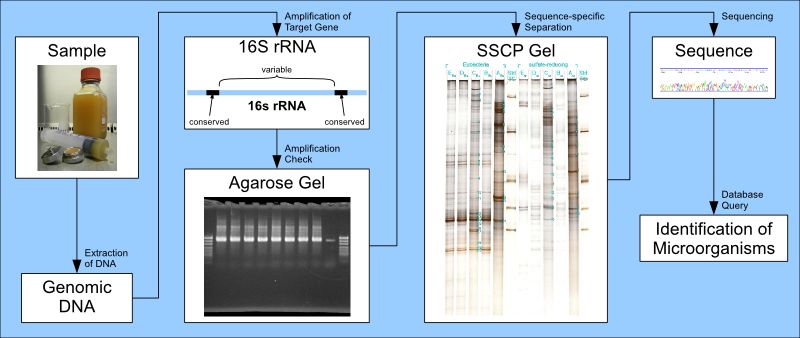
Screening
In the first step
- the DNA of all organisms in the sample material is extracted
- the microbial target DNA is universally amplified
- the amplified, double-stranded DNA gets controlled on an agarose gel
- one of the DNA strands gets removed
- the so-formed single strands undergo a sequence-specific separation on a gel
- the gel with the separated DNA gets stained
According to this scheme a gel is generated in which every lane contains the single-stranded DNA from a sample. DNA fragments with identical sequence move at the same speed and form a band. Different bands contain DNA with a different sequences. Which, if the target sequence was selected accordingly, relate to different species.
This gel allows first judgements regarding the amount and diversity of microorganisms present in a sample. This is why the customer gets this gel picture to select the bands (= species) of interest.
Identification
In a second step the DNA from the customer-selected bands gets extracted and sequenced. These sequences are compared with sequence databases, comprising sequences of known microorganisms. This database query results in the next-homologous sequence and species
AMODIA develops not only customer-specific systems for different target-genes. We also create and maintain customer-specific sequence databases. Get in contact!


